A comprehensive study of the effects of chromosomal rearrangements, in monkeyflowers
A study of the effect of chromosomal inversions on gene expression.
Chromosomal inversions are known to have strong phenotypic effects, for example in determining life histories, in many organisms. However the quantitative importance of chromosomal rearrangements, which include inversions and translocations, is not known, because only rearrangements of known effects have been historically studied.
We aim to advance our understanding of the role of chromosomal rearrangements in evolution by studying genome-wide inversions at the level of the individual, the population and the species.
To do so, we are sequencing and assembling multiple genomes at varying degrees of phylogenetic distance from each other in order to characterise the accumulation of chromosomal rearrangements over evolutionary time. We are simultaneously obtaining full transcriptome data from the same individuals to quantify the chromosomal rearrangement effects in gene expression.
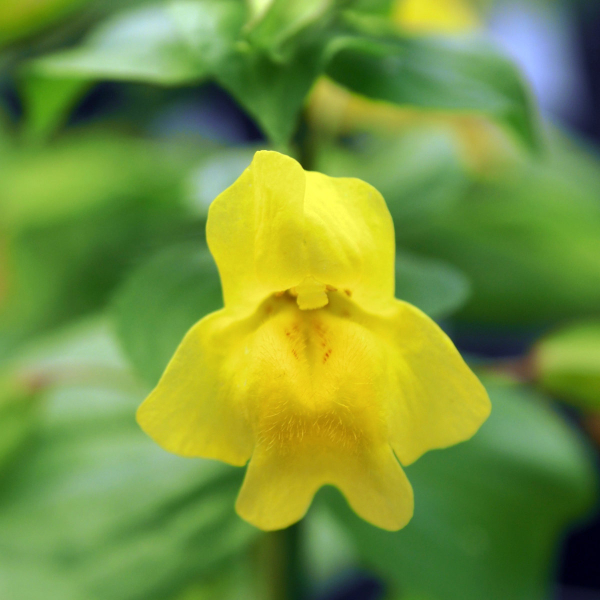

The organism of choice is the Mimulus species complex, because it harbours known inversions with phenotypic effects. In addition, Mimulus species have small genomes and high rates of divergence between populations and species, while retaining genetic compatibility, making them ideal to study chromosomal rearrangements.
This project is supported by a National Science Foundation grant to John Kelly.